Introducing Illumina Complete Long Read sequencing technology
This novel technology will bring more light to even the darkest corners of the genome. Illumina Complete Long Reads helps resolve the most challenging regions of the genome and makes long-read sequencing accessible and streamlined by enabling short and long reads from a single platform.
Illumina Complete Long Read technology will power a portfolio of novel, high-performance assays using standard next-generation sequencing (NGS) workflows and trusted Illumina sequencing by synthesis (SBS) chemistry.
Key features
Accessible and cost-effective
Both long and short reads from your NovaSeq platform. Easily integrates into standard NGS workstreams with included cloud analysis on BaseSpace Sequence Hub.
Robust and flexible
Low (50 ng) DNA input, tolerant to contaminants and freeze/thaw cycles. Generate contiguous long-read sequences with N50 of 5-7kb with as low as 5 ng.
Scalable and streamlined workflow
Simple one-day library prep workflow without specialized equipment. Assay 4 samples per run, up to 3000 WGS samples per year, with NovaSeq X Series.
Highly accurate
Powered by proven Illumina SBS and DRAGEN analysis on BaseSpace Sequence Hub. Highly accurate (99.87%) as measured by PrecisionFDA Challenge v2.
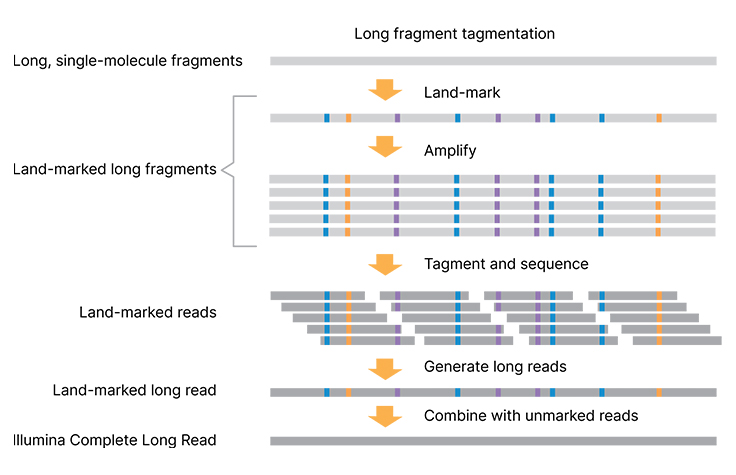
How the Illumina Complete Long Read technology works
Tagmentation is used to normalize long fragment sizes. Long fragments are “land-marked” to capture single-molecule, long-read information and amplified. Marked fragments are tagmented to standard libraries for sequencing. Marked and unmarked data are combined to generate highly accurate complete long reads.
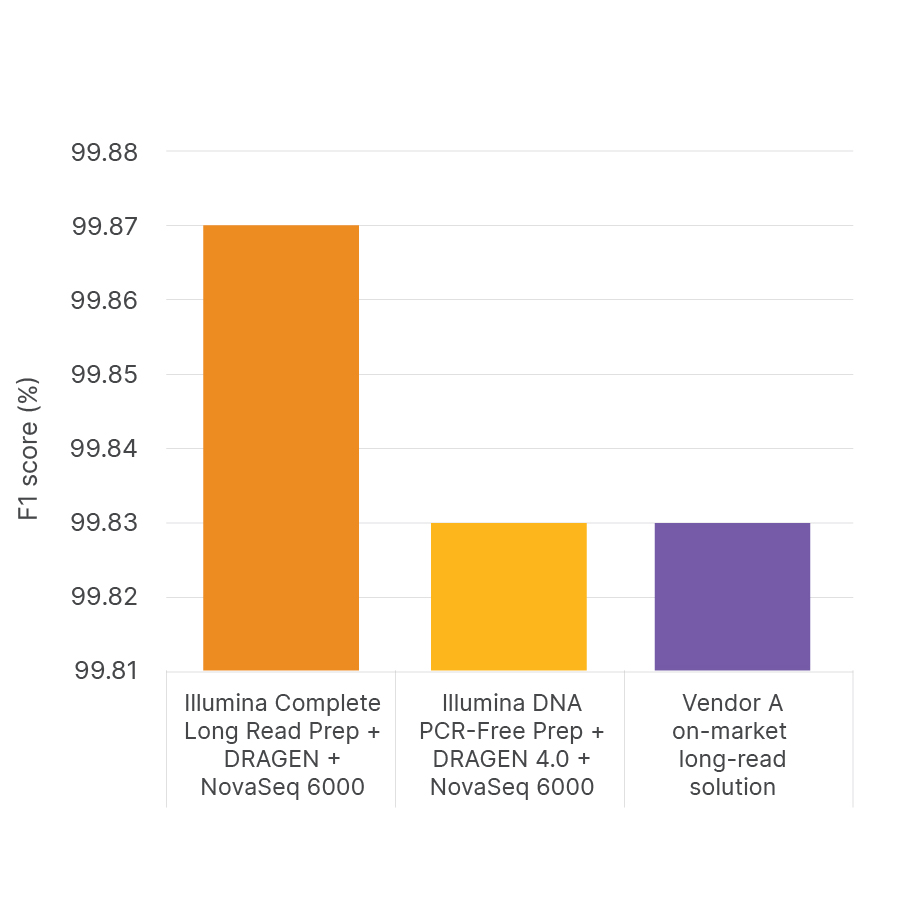
Illumina Complete Long Reads and PrecisionFDA Truth Challenge V2 data sets
A new standard for accuracy
Illumina Complete Long Read Prep, Human delivers unprecedented accuracy for variant calling with PrecisionFDA Truth Challenge v2 data sets,1 as measured by F1 score (%), a calculation of true positive and true negative results as a proportion of total results.
- Improves upon the exceptional, award-winning accuracy of Illumina SBS + DRAGEN analysis1-3
- Can be used to augment existing WGS data sets or as a reflex tool for deeper variant discovery
- Delivers highly accurate, reliable results across samples of variable quality
Illumina Complete Long Read products
Illumina Complete Long Read Prep, Human
Illumina Complete Long Read Prep, Human, the first product based on Illumina Complete Long Reads, is designed for human whole-genome sequencing (WGS).
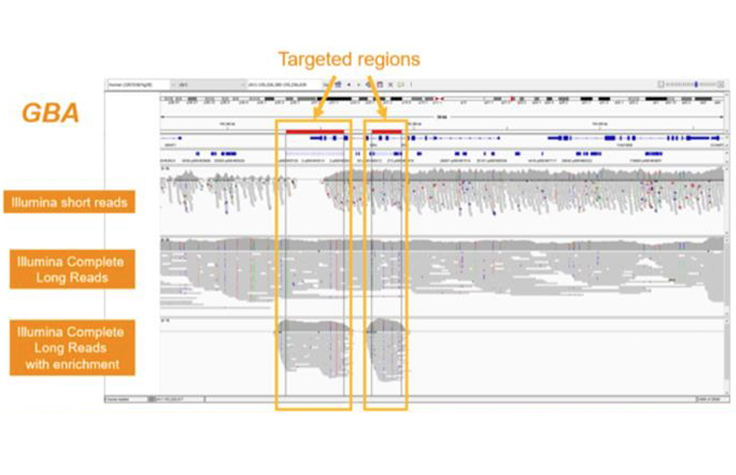
Illumina Complete Long Read Prep with Enrichment (Launching 2H2023)
In addition to the whole-genome assay, Illumina long-read technology is compatible with enrichment. Targeted solutions will focus on regions known to benefit from additional insight with longer reads. Future Illumina Complete Long Read products with enrichment can create additional flexibility and scalability for cost-effective human WGS.
Illumina Complete Long Read technology
Featured workflow
Extract
Extract DNA from blood or saliva with no specialized protocols, no shearing, and no size selection required
Prepare
Prepare libraries in a single day, with recommended 50 ng or as low as 10 ng input DNA, using standard lab equipment
Sequence
Sequence prepared libraries on the NovaSeq 6000 System or NovaSeq X Series
Analyze
Analyze using DRAGEN Illumina Complete Long Read app on BaseSpace Sequence Hub
Interested in the power of Illumina Complete Long Reads for your lab?
Speak with one of our specialists about how your workflow and analysis can benefit from the accuracy and scalability of our long-read technology. Fill out the form to request a quote for order today.Supporting data
Illumina Complete Long Read data can improve alignment and variant calling in traditionally challenging regions like repetitive regions, highly polymorphic regions, pseudogenes and paralogs, large insertion–deletion variants, and structural variants.
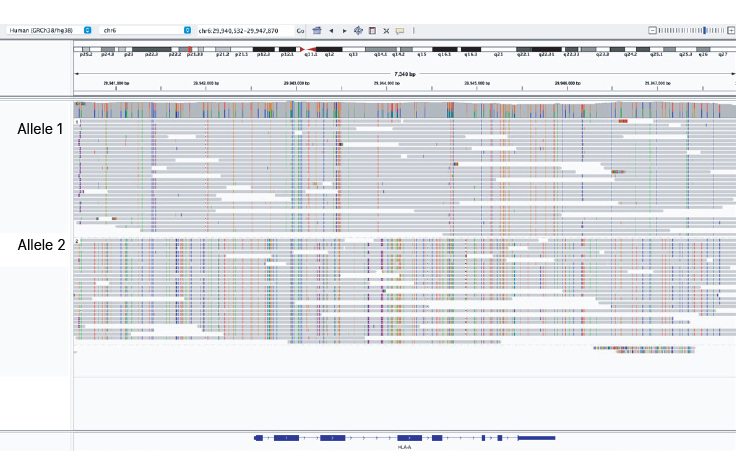
Long-read phased sequencing enables clear haplotype assignment
Illumina Complete Long Reads can resolve highly polymorphic regions like the HLA-A gene. Uniform coverage in the HLA region enables accurate phasing of HLA alleles.

Long-read sequencing can detect large deletions
A heterozygous 180 bp deletion in the SEMG1 gene is clearly detected by both Illumina Complete Long Reads and on-market long reads.
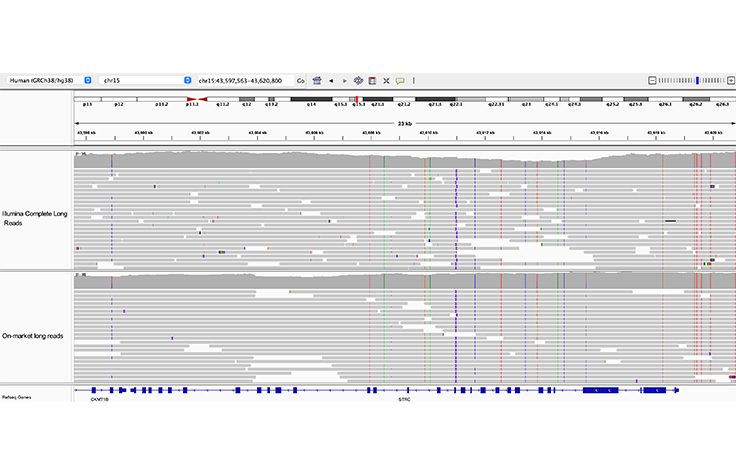
Long-read sequencing can resolve pseudogenes
Both Illumina Complete Long Reads and on-market long reads clearly resolve the 23 kb STRC gene from its pseudogene, pSTRC.
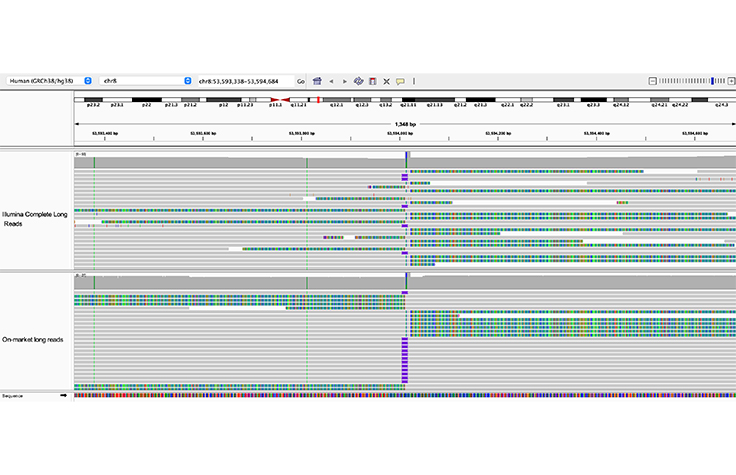
Long-read sequencing can identify STR insertions
Insertions in short tandem repeat (STR) regions are clearly detected by both Illumina Complete Long Reads and on-market long reads.

Long-read sequencing can resolve pseudogenes
Both Illumina Complete Long Reads and on-market long reads clearly resolve the SULT1A1 gene (a pharmacogenomic marker) from its pseudogene, SULT1A2.
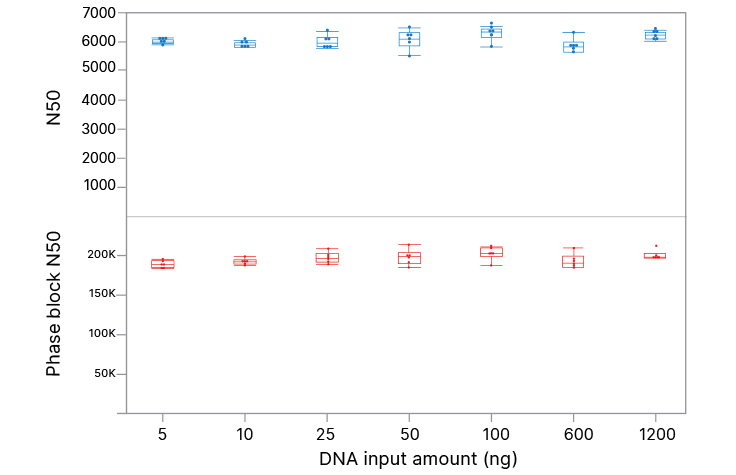
High-quality performance across a wide range of DNA input
Illumina Complete Long Read Prep, Human with DNA inputs from 5 ng to 1200 ng (in triplicate) generates similar data quality for N50 and phase block N50. N50 is defined as the sequence length of the shortest contig (or phase block) at 50% of the total assembly length.
Compatible with trusted Illumina sequencing systems
Illumina Complete Long Read Technology is compatible with the NovaSeq X Plus, NovaSeq X, and NovaSeq 6000 Sequencing Systems, giving you access to both long- and short-read data on the same instrument.

Resources
Illumina Complete Long Reads webinar
In this presentation, we share use cases of Complete Long Reads and highlight research being done by collaborators around the world.
Long-read technology
This recorded presentation from our ASHG 2022 CoLab talk describes the benefits of our novel long-read technology and how it is complemented by other technical innovations in our roadmap.
Illumina Long Read technology performance data comparison
Illumina Complete Long Read technology will help researchers address the edges of the genome that are the most challenging to map.
References
- PrecisionFDA. Truth Challenge V2: Calling Variants from Short and Long Reads in Difficult-to-Map Regions. precision.fda.gov/challenges/10
- Mehio R, Ruehle M, Catreux S, et al. DRAGEN Wins at PrecisionFDA Truth Challenge V2 Showcase Accuracy Gains from Alt-aware Mapping and Graph Reference Genomes. illumina.com/science/genomics-research/dragen-wins-precisionfdachallenge-showcase-accuracy-gains.html
- Illumina. Accuracy improvements in germline small variant calling with the DRAGEN Bio-IT Platform. illumina.com/content/dam/illumina/gcs/assembled-assets/marketing-literature/dragen-v4-accuracy-app-note-m-gl-01016/dragen-v4-accuracy-app-note-m-gl-01016.pdf
